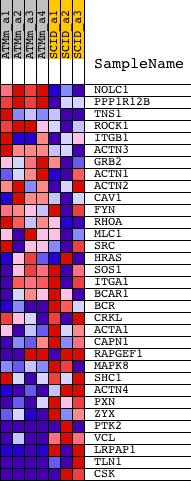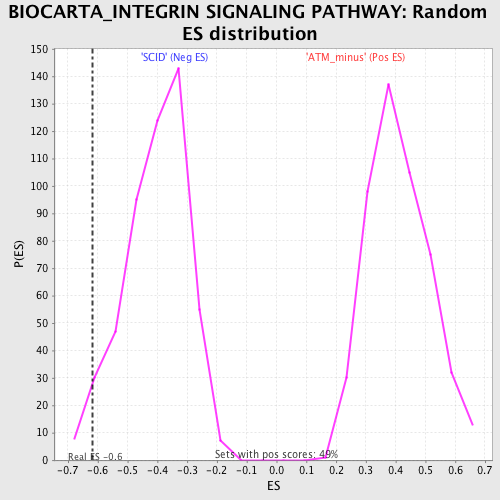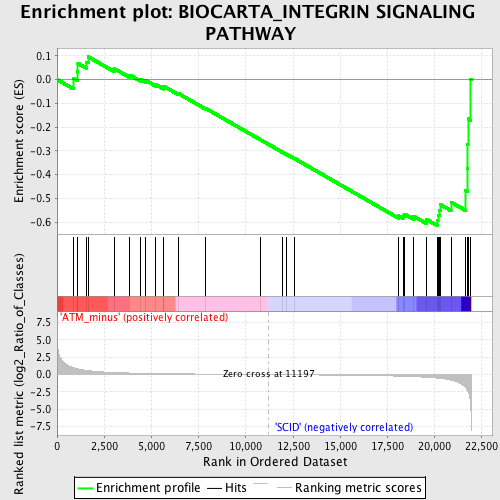
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_SCID.phenotype_ATM_minus_versus_SCID.cls #ATM_minus_versus_SCID |
| Phenotype | phenotype_ATM_minus_versus_SCID.cls#ATM_minus_versus_SCID |
| Upregulated in class | SCID |
| GeneSet | BIOCARTA_INTEGRIN SIGNALING PATHWAY |
| Enrichment Score (ES) | -0.61672884 |
| Normalized Enrichment Score (NES) | -1.5359133 |
| Nominal p-value | 0.031434186 |
| FDR q-value | 0.9031631 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NOLC1 | 1428869_at 1428870_at 1450087_a_at | 853 | 0.995 | 0.0060 | No | ||
| 2 | PPP1R12B | 1431462_at 1434786_at 1441730_at 1457169_at 1457338_at 1460091_at | 1082 | 0.823 | 0.0328 | No | ||
| 3 | TNS1 | 1419283_s_at 1428650_at 1449405_at | 1103 | 0.808 | 0.0684 | No | ||
| 4 | ROCK1 | 1423444_at 1423445_at 1441162_at 1446518_at 1450994_at 1460729_at | 1574 | 0.592 | 0.0737 | No | ||
| 5 | ITGB1 | 1426918_at 1426919_at 1426920_x_at 1427771_x_at 1438119_at 1452545_a_at | 1653 | 0.567 | 0.0958 | No | ||
| 6 | ACTN3 | 1418677_at | 3049 | 0.293 | 0.0453 | No | ||
| 7 | GRB2 | 1418508_a_at 1449111_a_at | 3849 | 0.223 | 0.0189 | No | ||
| 8 | ACTN1 | 1427385_s_at 1428585_at 1452415_at | 4431 | 0.186 | 0.0008 | No | ||
| 9 | ACTN2 | 1448327_at 1456968_at | 4698 | 0.172 | -0.0036 | No | ||
| 10 | CAV1 | 1449145_a_at | 5222 | 0.149 | -0.0207 | No | ||
| 11 | FYN | 1417558_at 1441647_at 1448765_at | 5640 | 0.133 | -0.0338 | No | ||
| 12 | RHOA | 1437628_s_at | 5643 | 0.133 | -0.0278 | No | ||
| 13 | MLC1 | 1448139_at | 6446 | 0.108 | -0.0596 | No | ||
| 14 | SRC | 1423240_at 1450918_s_at | 7866 | 0.070 | -0.1212 | No | ||
| 15 | HRAS | 1422407_s_at 1424132_at | 10749 | 0.009 | -0.2523 | No | ||
| 16 | SOS1 | 1421884_at 1421885_at 1421886_at | 11915 | -0.015 | -0.3048 | No | ||
| 17 | ITGA1 | 1439713_at 1446569_at 1455251_at | 12165 | -0.020 | -0.3153 | No | ||
| 18 | BCAR1 | 1439388_s_at 1450622_at | 12572 | -0.029 | -0.3325 | No | ||
| 19 | BCR | 1427265_at 1452368_at 1455564_at | 18075 | -0.220 | -0.5737 | No | ||
| 20 | CRKL | 1421953_at 1421954_at 1425604_at 1436950_at | 18321 | -0.238 | -0.5742 | No | ||
| 21 | ACTA1 | 1427735_a_at | 18396 | -0.245 | -0.5664 | No | ||
| 22 | CAPN1 | 1417228_at 1417229_at | 18878 | -0.290 | -0.5753 | No | ||
| 23 | RAPGEF1 | 1421146_at 1427006_at 1441386_at | 19549 | -0.382 | -0.5886 | No | ||
| 24 | MAPK8 | 1420931_at 1420932_at 1437045_at 1440856_at 1457936_at | 20166 | -0.512 | -0.5936 | Yes | ||
| 25 | SHC1 | 1422853_at 1422854_at | 20174 | -0.514 | -0.5707 | Yes | ||
| 26 | ACTN4 | 1423449_a_at 1444258_at | 20263 | -0.543 | -0.5501 | Yes | ||
| 27 | PXN | 1424027_at 1426085_a_at 1456135_s_at | 20305 | -0.553 | -0.5270 | Yes | ||
| 28 | ZYX | 1417240_at 1438552_x_at | 20865 | -0.796 | -0.5165 | Yes | ||
| 29 | PTK2 | 1423059_at 1430827_a_at 1439198_at 1440082_at 1441475_at 1443384_at 1445137_at | 21633 | -1.852 | -0.4679 | Yes | ||
| 30 | VCL | 1416156_at 1416157_at 1445256_at | 21722 | -2.191 | -0.3729 | Yes | ||
| 31 | LRPAP1 | 1426696_at 1426697_a_at 1436609_a_at 1452148_at | 21730 | -2.234 | -0.2722 | Yes | ||
| 32 | TLN1 | 1436042_at 1448402_at 1457782_at | 21764 | -2.390 | -0.1657 | Yes | ||
| 33 | CSK | 1423518_at 1439744_at | 21885 | -3.839 | 0.0023 | Yes |